« There are three kinds of lies: lies, damned lies, and statistics ». This sentence by British Prime Minister Benjamin Disraeli originates from an inappropriate use of statistical tools to support an intuition or conviction. Although censurable, this practice unfortunately has its counterpart in research: p-value hacking. It consists in dredging the data until the desired significance is obtained for a hypothesis tested, to the detriment of some statistical rules that are violated. In most cases, this is done in good faith, since the objective is to get the best from the data while complying with publication requirements. Nevertheless, it is essential to limit this practice, which leads to many false discoveries.
This is practically the case in proteomics, due to the intrinsic proteome complexity, but also to the rapid evolution of analytical technologies. This is one of the reasons why many bioinformatics and biostatistics tools regularly blossom in the literature
[1], with the promise of overcoming the triple limit of big proteomics data: their large size, their big dimensionality, and their great complexity. However, the simplicity of these tools cannot hide the need for a minimum of theoretical understanding to use them correctly.
It is with this objective in mind that researchers at IRIG have made a special effort in recent years to disseminate good practices in data science for proteomics
[2-4]. They published an introduction to FDR theory (False Discovery rate, an ubiquitous quality control measure), they disambiguated a set of terms with different meanings in artificial intelligence and analytical chemistry, and also proposed five steps to improve the quality control of differential proteomic analysis between several samples.
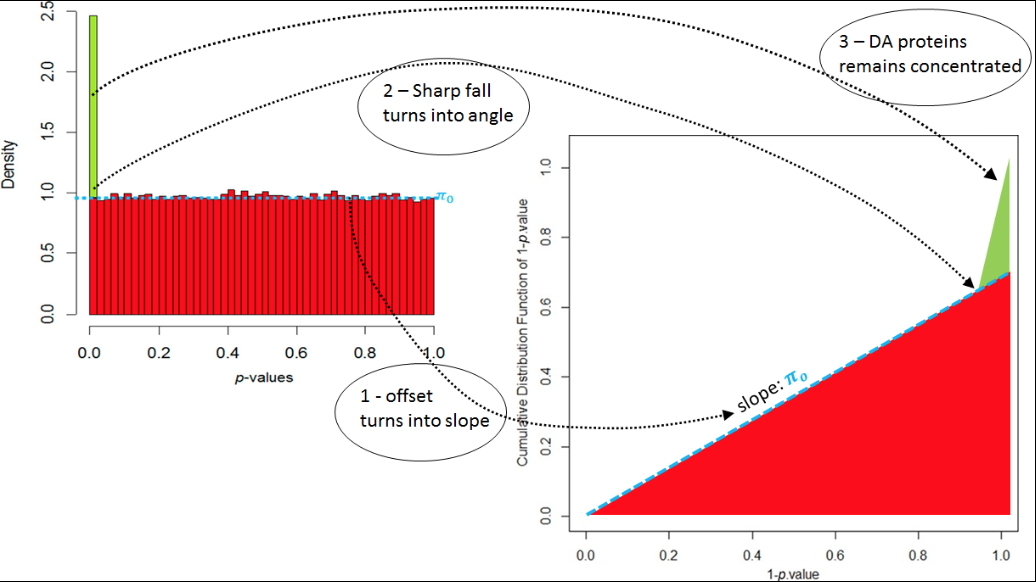
Graphical construction allowing to visually estimate the quality of the p-values calibration.