Biological membranes are organized as two layers of lipids in which proteins are partially or totally embedded. The lipid composition of the two leaflets is different, thereby controlling many biological processes such as membrane trafficking or cell signaling. This lipid asymmetry results from many mechanisms whose main contributors are transmembrane lipid transport proteins. Floppases catalyse the transport of lipids from the inner (cytosolic) to the outer (exoplasmic) membrane leaflet and flippases do the opposite. In humans, mutations of several flippase homologs are involved in rare forms of intrahepatic cholestasis or neurological disease (cerebellar ataxia syndrome associated with intellectual disability). Studies in rodent show flippases are essential for insulin secretion, the shape of the red blood cells or the survival of the photoreceptors on retina. In plants, one of the orthologue is involved in heat stress responses. Significant efforts are made to determine the structure of these proteins and thus to clarify the lipid transport mechanisms. Flippases are the only subtype of P-ATPases to transport lipid. The other ones transport either ions or very small molecules compared to aminophospholipids. Regarding the high conservation of residues and putative structures, how have P4-ATPases evolved to provide a pathway of sufficient size to allow for translocation of lipids instead of ions ? This classical conundrum posed by the nature of the substrate is refered to as ‘giant substrate” problem.
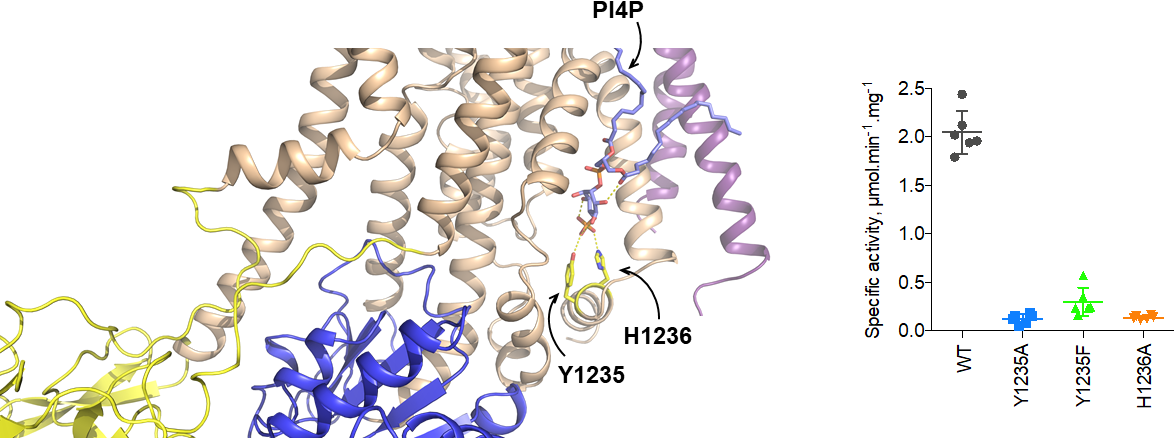
Teams from I2BC (UPSud/CEA/CNRS), University of Aarhus and Max Planck Institute of Biophysics in Franckfurt have collaborated to determine the structure of the Drs2p-Cdc50p flippase from S. cerevisiae by cryo-electron microscopy. The C-terminal tail of Drs2 contains an auto-inhibitory domain. Binding of PI4P[1] reduces partially this auto-inhibition. This conformation is called intermediate. C-terminal tail truncation makes the enzyme active in presence of PIP4. The structure of this flippase was studied in three different conformations: a conformation in which it is auto-inhibited, a conformation in which it is fully active and a third intermediate conformation. This first for the flippase family was made possible by the cryo-electron microscopy (imaging technique for wich three researchers won the Nobel Prize for chemistry in 2017). The three structures obtained at resolutions of 2.8 to 3.7 Å reveal the mechanism by which the C-terminal end of Drs2p inhibits complex activity and the first steps in auto-inhibition relief by PI4P. Comparison of the three structures also reveals the existence of a cavity through which the polar headgroup of the substrate lipid (mainly phosphatidylserine) might be transported, as it travels from one layer of the cell membrane to the other.
Des équipes de l’Institut de biologie intégrative de la cellule (Université Paris-Sud/CEA/CNRS), de l’Université d’Aarhus et de l’Institut Max Planck à Francfort se sont associées pour déterminer la structure de la protéine Drs2 en complexe avec sa sous-unité associée Cdc50, une flippase de la levure. Drs2 contient un segment à son extrémité C-terminale qui permet l’auto-inhibition de son activité. Cette auto-inhibition est partiellement levée en présence de phosphatidylinositol-4-phosphate (PI4P)[2] ; on parle alors de conformation intermédiaire. La troncation de l’extrémité C-terminale, en présence de PI4P, rend l’enzyme pleinement active. Les équipes ont obtenu la structure du complexe Drs2-Cdc50 dans chacune de ces trois conformations : auto-inhibée, intermédiaire et active. Cette première pour la famille des flippases a pu se faire grâce à la cryo-microscopie électronique (technique d’imagerie qui a valu le prix Nobel de chimie à trois chercheurs en 2017). Les trois structures, obtenues à des résolutions de 2,8 à 3,7 Å, révèlent le mécanisme par lequel l’extrémité C-terminale de Drs2 inhibe l’activité du complexe et les premières étapes de la levée de cette auto-inhibition par le PI4P (figure 1). La comparaison des structures révèle également l’existence d’une cavité par laquelle les chercheurs proposent que la tête polaire du lipide substrat (la phosphatidylsérine principalement) soit prise en charge lors de son cheminement d’un feuillet à l’autre des membranes cellulaires (figure 2).
|
|
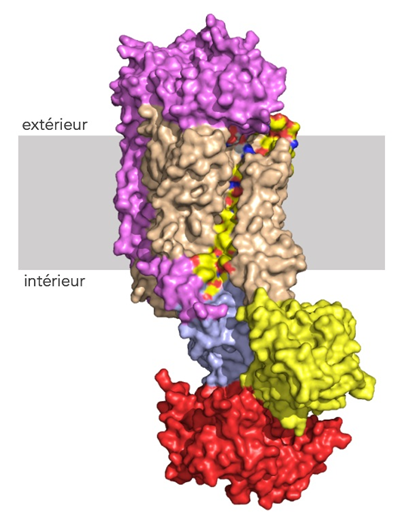
Possible lipid pathway through membrane bilayer. Transmembrane helix 4, along the transmembrane cavity of Drs2, is labeled in yellow. The rest of the transmembrane domain is in pale tane brown.Cytosolic domain A, N and P are respectively yellow, red and light blue. Cdc50 is iink. © G. Lenoir/I2BC
|
This work describes for the first time the molecular architecture of a flippase. Other structures of P4-type ATPases, in the presence of their lipid substrate and / or regulatory partners, must be obtained to fully elucidate the mechanism of lipid « flip » and see if it is conserved from one eukaryote to another.
These results have also been shared trough a press release.
[1]
Phosphatidylinositol-4-phosphate (PI4P) is a phospholipid abundant in Golgi membranes.