In most organisms there are repeated and mobile DNA sequences that are capable of moving within their genome. Where these transposable elements (TEs) fit into the DNA determines their impact. New insertions can contribute to the development of new cellular functions and thus to the long-term adaptation of organisms to different environments, but they can also pose a threat to the integrity of genomes by creating deleterious mutations. In humans, more than a hundred hereditary diseases have been attributed to
de novo insertions of TEs. The distribution of TEs in the genome is rarely random and is the result of a balance between two processes. On the one hand, selection leads to the elimination of highly damaging insertions and the maintenance of beneficial ones. On the other hand, TEs have evolved and put in place mechanisms that direct their integration into "safe" places in the genome, where their insertions will have a minimal negative effect on the organism.
Researchers at IRSL and I2BC have been interested for several years in understanding the mechanisms behind the targeted integration of TEs into the genome. Their studies focus on Ty1, a retrotransposon[1] of the yeast
Saccharomyces cerevisiae, which integrates preferentially upstream of the transcribed genes by an enzyme complex, RNA polymerase III, specialized in the synthesis of short RNAs. A previous study had shown that the targeted integration of Ty1 is dependent on the interaction between two proteins: IN1, the integrase encoded by Ty1 itself, and AC40, a subunit common to the RNA polymerase III and RNA polymerase I complexes.
In a new study, published in
the EMBO Journal, the researchers identify the short sequence of IN1 that interacts with AC40. The interaction between this domain of IN1 and AC40 allows the recruitment of IN1 on genes transcribed by RNA polymerases I and III. On the other hand, it allows the integration of Ty1 only upstream of the genes transcribed by RNA polymerase III. The introduction of this sequence into the integrase of the retrotransposon Ty5 - which is normally inserted in the subtelomeric regions - directs the integration of Ty5, like that of Ty1, upstream of the genes transcribed by RNA polymerase III.
Beyond the breakthrough for basic research, this study could help improve retroviral vectors used in gene therapy to transfer genes within cells. These vectors are often integrated into gene-rich regions where they can have mutagenic effects. New retroviral vectors could be designed based on the interaction between IN1 and AC40 to target their upstream integration of genes transcribed by RNA polymerase III and potentially limit mutagenic risks.
Contact for CEA-Joliot:
Joël Acker (joel.acker@cea.fr)
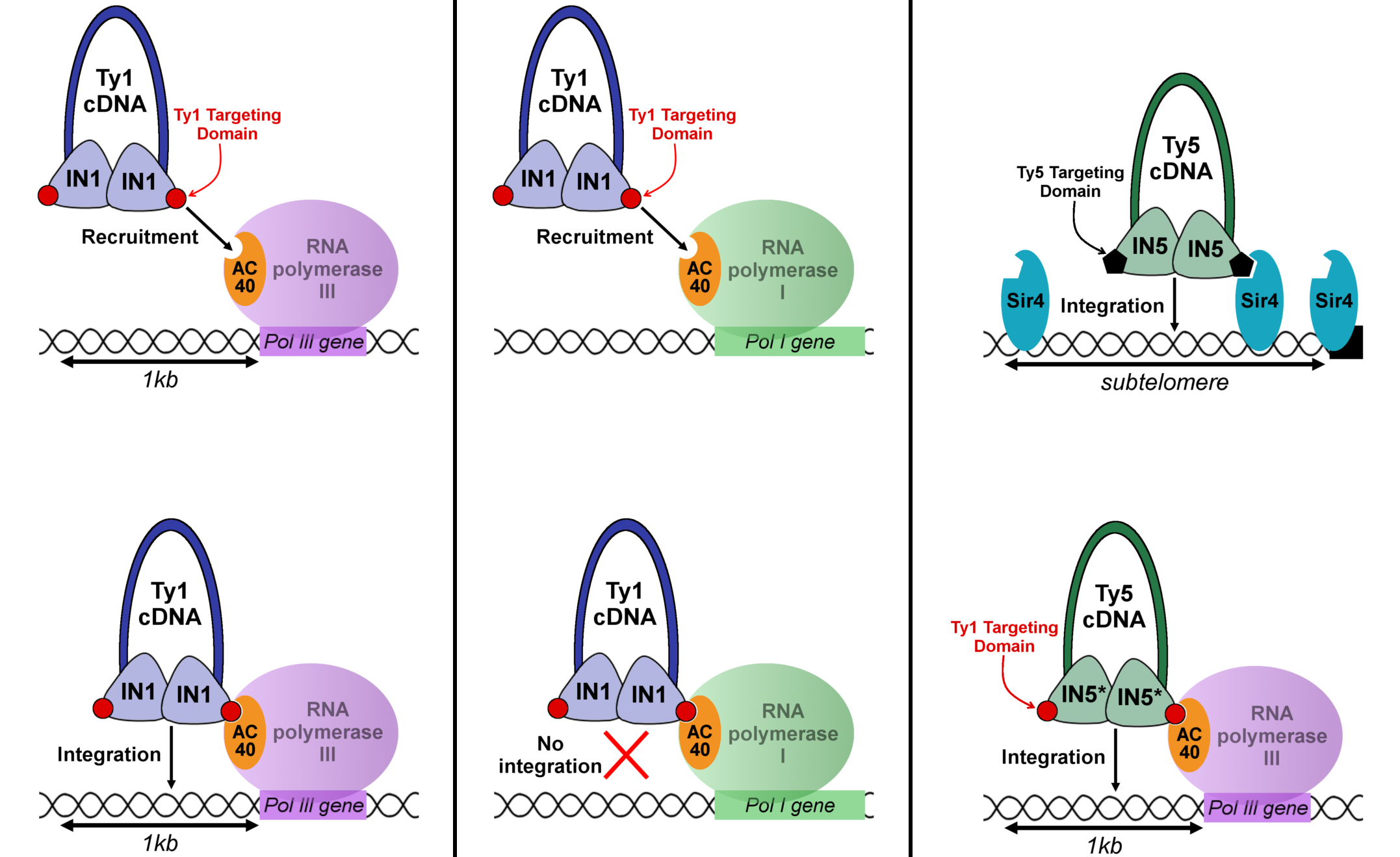
A short sequence of IN1 interacts with AC40, the common subunit of the Pol I and Pol III RNA polymerases. This interaction directs IN1 recruitment to all genes transcribed by Pol I and Pol III. It targets the integration of Ty1 only to the genes transcribed by Pol III. The replacement of the Ty5 retrotransposon integrase targeting domain by this motif redirects Ty5 integration from the telomeric regions to those upstream of the Pol III transcribed genes. © A. Asif-Laidin et al.
[1] Retrotransposons replicate through an RNA.