Staphylococcal enterotoxins (SEs), produced by the bacteria Staphylococcus aureus, are responsible for around 10% of food poisoning outbreaks worldwide. In this case, the gastrointestinal symptoms observed are linked to the ingestion of SEs, heat-resistant proteins, produced by the bacteria which could develop in the food, following contamination of the raw material used or during the process of elaboration.
SEs constitute a very diverse family of proteins: 27 types of toxins are known today, each with many genetic variants. If food poisoning by these proteins is suspected, they are looked for in the offending foods or food ingredients in order to remove those contaminated from the food chain. To avoid false negatives during these analyzes, it is necessary to be able to detect as many different SEs as possible. This identification is now based on the immuno-analysis of suspicious samples, a method with high sensitivity but possibly influenced by the nature of the food matrices, or even on PCR, which highlights the presence of genes encoding SEs, but which does not provide direct evidence for the presence of these toxins. Mass spectrometry is also used but with poor coverage of SEs diversity. It is therefore necessary to develop new methods capable of identifying this diversity in different food matrices.
To meet this challenge, researchers from SPI (DMTS), in collaboration with ANSES, used targeted high-resolution mass spectrometry (MS). By carefully considering the amino acid variations within each type of SE and the sequence homology between the different types, 3 to 8 proteotypic peptides from each of the 8 most frequent SEs were selected to be searched for by targeted MS in the samples. The sensitivity of the assay was increased, thanks to a first step of enrichment by immuno-capture and its multiplexing capacity was improved by a thorough optimization of sample preparation and MS acquisition protocols. The assay was then validated in dairy products, and the quantification of SEs successfully demonstrated in real samples, collected during outbreaks of staphylococcal food poisoning. Finally, the ability of the method to detect various sequence variants has also been illustrated.
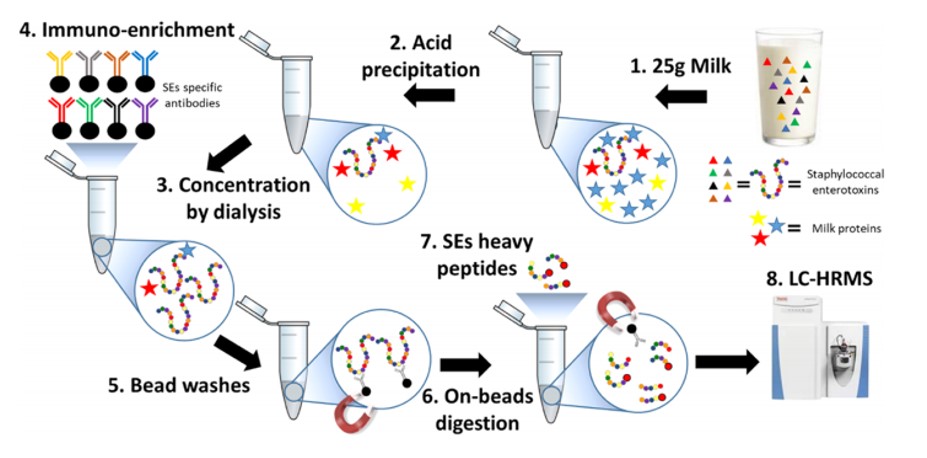
Diagram of the different steps of the test developed by the SPI for this study.
© J. Agric. Food Chem. 2021
In conclusion, a multiplex assay has been developed to simultaneously identify and quantify, in dairy samples, the eight most frequent SEs, at symptomatic concentrations. In the future, this method will have to be tested on other food matrices and on a greater diversity of toxin variants to confirm the value of this assay in food poisoning epidemics.
ANSES : agence nationale de sécurité sanitaire de l'alimentation, de l'environnement et du travail
Contact: François Becher francois.becher@cea.fr