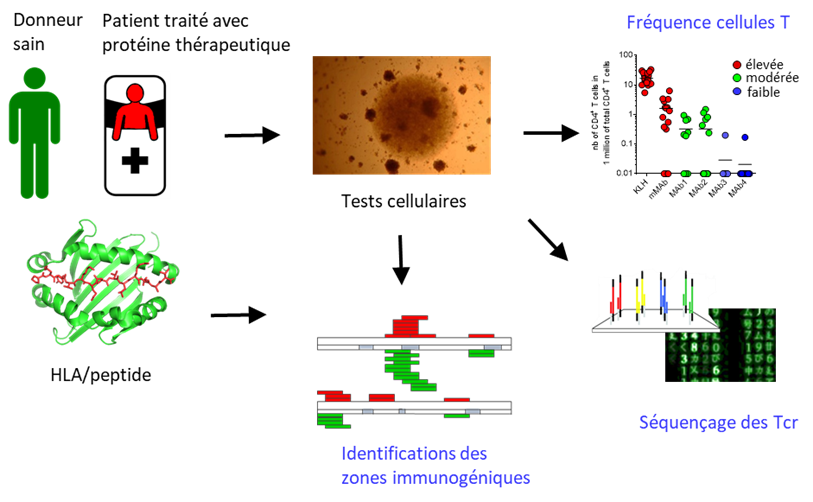
The Antibody Engineering team develops novel antibody-based ligands for a range of applications, including therapeutic, imaging, and diagnostic purposes. It focuses particularly on designing high-affinity antibodies with controlled selectivity and low immunogenicity, enabling administration in humans while minimizing the risk of immune responses. The team has developed protein molecular engineering tools that combine a high-throughput screening method, Yeast Surface Display, with high-throughput NGS sequencing technologies. In particular, it implements a Deep Mutational Scanning (DMS) approach aimed at identifying, within protein sequences, the key amino acids that play a functional role in the property of interest. DMS data notably help elucidate protein–protein interfaces (such as epitopes or paratopes). They are also used to guide the engineering of optimized molecules, particularly for affinity maturation or antibody selectivity engineering.
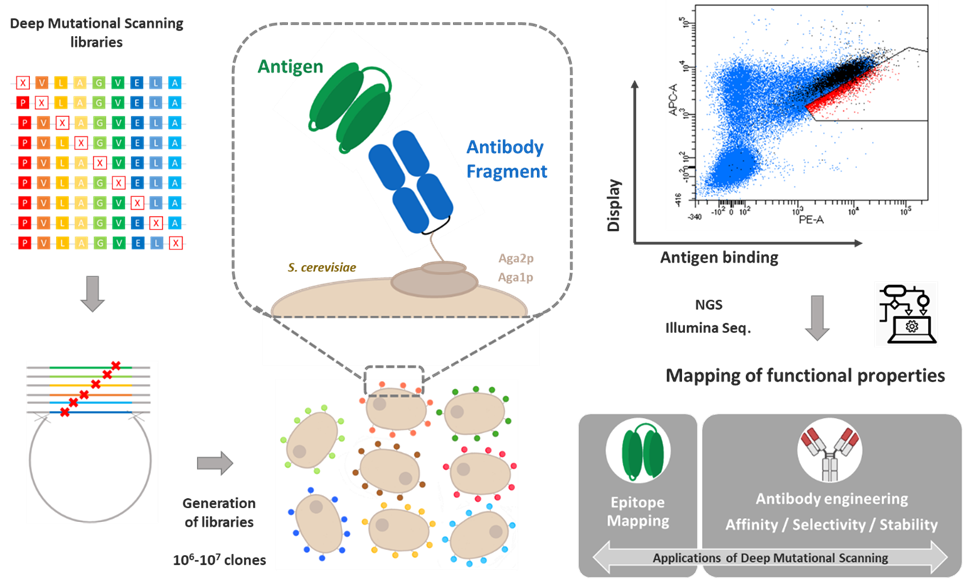
These technologies led to the creation of the antibody engineering service company Deeptope (http://deeptope.com).
We also conduct activities aimed at discovering novel VHH ligands from both synthetic libraries and immunization-derived libraries, in order to identify antibodies with optimized functional characteristics. In parallel, we develop artificial intelligence–based approaches designed to perform differential screening of antibodies according to their antigen-binding interface. These strategies, enable the selection of antibodies targeting specific epitopes and the achievement of precisely characterized and controlled modes of action. This work is carried out in the context of applications in oncology, in the fight against various emerging infectious diseases, as well as for agents of the CBRN-E domain.
Synthetic biology and evolution
The “Synthetic Biology and Evolution" (SBE) team combines biomolecular engineering, synthetic biology, and generative artificial intelligence (AI) to design innovative biological systems and proteins. Our approach integrates high-throughput methods (multiplexed assays of variant effects, MAVE; NGS) and proprietary technologies such as in vitro DNA assembly (SLiCESBE) and bacterial genome engineering (bDICESBE). A core technology underpinning its activities is the qB2HSBE system (quantitative bacterial two-hybrid), which generates high-quality protein–protein interaction data ideally suited for AI model training and for accelerating multispecific antibody engineering.
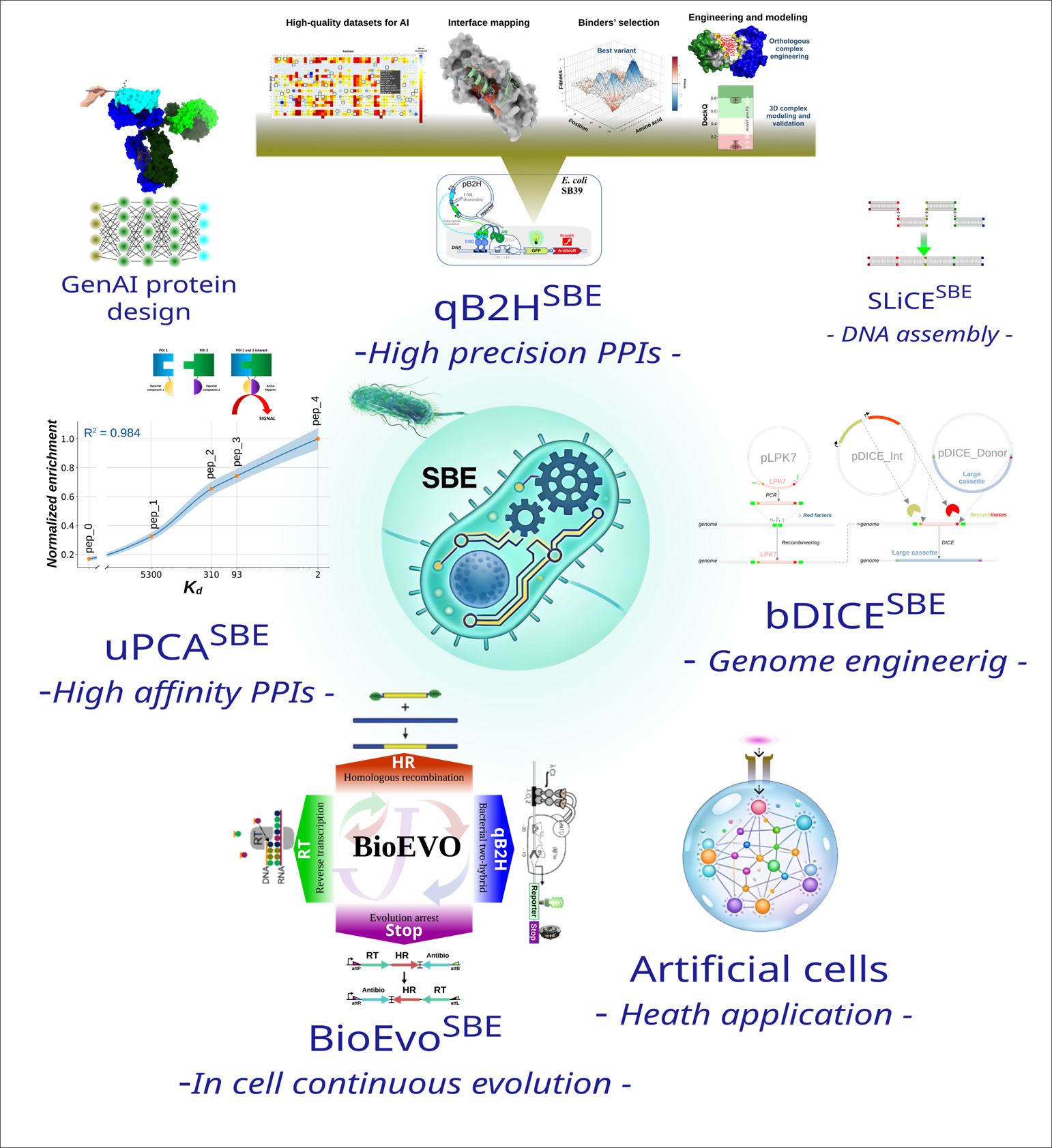
In parallel, our generative AI pipelines model antigen–antibody complexes at high resolution based on experimental data and design de novo proteins.We also work on:
- Programmable artificial cells dedicated to healthcare applications.
- uPCASBE: a universal protein complementation assay adaptable to diverse biological systems and offering a broad dynamic range.
- BioEvoSBE: a continuous molecular evolution system.
These research axes form a synergistic technological ecosystem, opening new perspectives in bioproduction, immunotherapy, and synthetic biology. At the interface between wet lab and dry lab, our tools democratize biological engineering through integrated solutions tailored to different scales.
Immunogenicity of proteins and antibodies
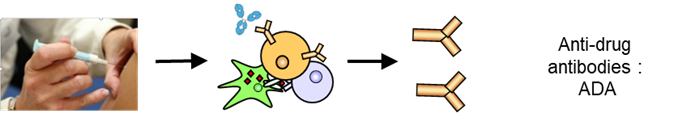
Protein immunogenicity results in the production, in treated patients, of antibodies directed against the protein (anti-drug antibodies: ADA).
Therapeutic antibodies and proteins offer many advantages, such as high selectivity and low toxicity, but they also present the drawback of being potentially immunogenic, meaning that they can trigger an immune response against themselves. The antibodies produced by patients and directed against a therapeutic protein may decrease or increase the pharmacokinetics of the protein, neutralize its therapeutic activity, or induce allergic or autoimmune symptoms.
Given the central role of CD4 T lymphocytes in initiating and regulating immune responses, assessing CD4 T-cell responses specific to therapeutic molecules enables us to better understand the origin of therapeutic protein immunogenicity. Our studies focus on identifying the sequences recognized by these cells (T-cell epitopes), characterizing the clonality of T cell repertoires specific to therapeutic proteins, defining the effector or regulatory phenotype of CD4 T cells specific to therapeutic proteins, and developing strategies to generate de-immunized molecules, i.e., molecules that no longer induce immune responses.
We also develop novel T-cell characterization tools based on T-cell receptor (TCR) cloning.