Understanding cellular processes at the atomic level requires the determination of the three dimensional structure of protein complexes and the characterization of their assembly/disassembly mechanisms. The laboratory is particularly interested in the description of the protein-protein interaction networks controlling genome stability : DNA damage signaling and repair, telomere architecture, nuclear envelope assembly and structure.
 Contacts
Contacts
DNA damage signaling and repair
Jean-Baptiste CHARBONNIER
jb.charbonnier@cea.fr
Telomere architecture
Marie-Hélène LE DU
marie-helene.ledu@cea.fr
Nuclear envelope assembly and structure
Sophie ZINN-JUSTIN
sophie.zinn@cea.fr
Control of genome stability
The team focuses on the description of complexes involved in the control of genome stability using an integrative structural biology approach: Small Angle X-ray Scattering (SAXS), Nuclear Magnetic Resonance (NMR), X-ray crystallography and electron microscopy provide experimental data, which are incorporated into molecular modeling calculations to obtain three-dimensional models of the complexes. Analysis of these models enables a better understanding of how protein-protein interaction networks control genome stability.
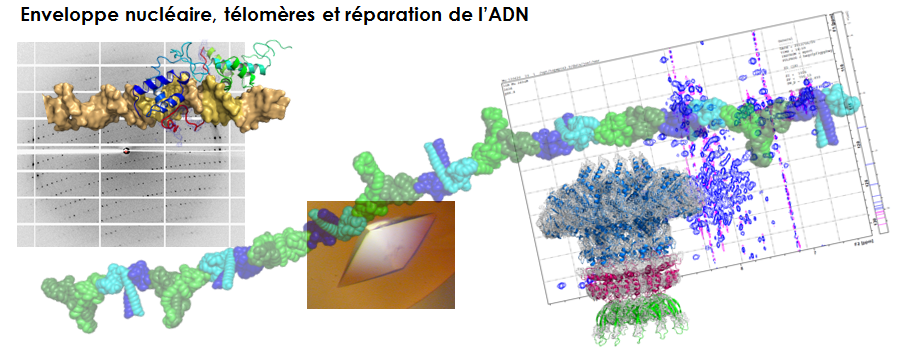
DNA damage signaling and repair
Because these networks are highly regulated and are interconnected, the protein complexes have a dynamical behavior: their composition and molecular architecture evolve all along their action in the cell. Furthermore, most of the proteins involved in these networks consist of several structural domains having distinct biological functions and interconnected by unstructured regions. This facilitates the coordination of their various biological functions. The laboratory aims at identifying high and low affinity interactions between protein domains, characterizing their mode of interaction with an atomic resolution and elucidating their mode of regulation using biochemical methods combined with fluorescence, calorimetry and Nuclear Magnetic Resonance. The impact of post-translational modifications on the protein three-dimensional structure and binding properties is investigated by Nuclear Magnetic Resonance
in vitro and in cell extracts. The presence of these complexes in the cell and their functional role are systematically addressed through collaboration with cell biologists working on DNA damage signaling and repair and nuclear organization.
The team includes a molecular modelling activity for structure determination of large protein complexes end the simulation of their dynamics in solution. The molecular modelling activity is based on various experimental data produced in the lab or obtained via collaborations (NMR, SAXS, EM, X–ray diffraction…). Simulation work aims at describing the molecular dynamics of the proteins at the atomic scale and the interaction thermodynamics of the proteins involved in these complexes. Two different approaches are used for molecular simulations: a standard well established method relying on the class I additive force field charmm22 (CHARMM/NAMD) and a more advanced methods currently developed in the lab (POLARIS(MD)) that relies on the non-additive polarizable class III force-field, TCPEp. The original approaches implemented in POLARIS(MD) and the optimization of the code for massively parallel computers achieved in collaboration with the INTEL/CEA/UVSQ Exascale Computing Research Laboratory allows considering simulations of molecular systems equivalent 105 to106 residues. We are also developing an efficient approach for identifying three-dimensional atom motif in the protein structure database (PDB) by systematic screening (STAMPS / site web RASMOT-3D, Debret et al., 2009;
http://biodev.cea.fr/rasmot3d/). This approach can be used to identify specific functions in proteins (Amrein et al., 2011) or for engineering (Tlatli et al., 2013).
Telomere architecture and Nuclear envelope assembly
Our team focuses on interactions regulating the response to DNA damage, the telomere architecture and the structure of the inner nuclear envelope. It has recently described the three-dimensional structure of the yeast endonuclease complex of the mismatch repair (Guéneau et al., 2013 NSMB) and of a protein filament involved in the ligation step of the main human DNA double-strand breaks repair pathway (Ropars et al., PNAS 2011). We addressed assembly of proteins involved in inhibition of DNA double-strand break repair at yeast (Matot et al, NAR 2012; Le Bihan et al, Acta Cryst D 2013) and human telomeres (Giraud-Panis et al, Front Oncol 2013). We finally got a first model of a complex involving a protein anchored at the inner nuclear envelope (Bourgeois et al., Science Signaling 2013). Deregulation of interactions involved in these processes is observed in aging-related diseases: cancers, progeroid syndromes. Our team collaborates with clinicians to understand the impact of mutations contributing to the onset of these diseases on the control of genome integrity and the structure of the nucleus.
Représentation schématique des protéines télomériques chez la levure Saccharomyces cerevisiae et chez les mammifères
L'assemblage des virus bactériens est un système très approprié pour étudier les mécanismes moléculaires impliqués dans la formation d'une machinerie macromoléculaire et sa fonction
Finally, the team is open to collaborations when integrative structural biology approaches can answer to biological questions. In particular, we have developed a strong partnership with the community of virologists working on bacteriophages, and we have determined the three-dimensional structure of several proteins belonging to the viral phage particles. This has led to the calculation of pseudo-atomic models of viral subcomplexes, based on nuclear magnetic resonance, crystallography X-ray and electron microscopy data (Lhuilier et al, PNAS 2009; Chagot et al, 2012 Proteins).
Expertises of the team scientist staff :
Jean-Baptiste CHARBONNIER : protein-protein interaction, crystallography, DNA damage repair
Philippe CUNIASSE : molecular modeling, integration of heterogeneous data, simulation
Pascal DREVET : production of proteins in insect cells, biochemistry, DNA damage repair
Bernard GILQUIN : molecular modeling, integration of heterogeneous data
Marie-Hélène LE DU : crystallography, SAXS, telomere
Michel MASELLA : molecular modeling, developments for simulation
Sophie ZINN-JUSTIN : Nuclear Magnetic Resonance, SAXS, nuclear envelope
Models in motion
Movie presenting a model of the complex between fragments of the transcription regulator Smad2 (in three different blue colors for the three subunits of the trimer) and the inner nuclear membrane protein MAN1 (in yellow, orange and red for the three MAN1 proteins, respectively)
The movie shows successively a superimposition of 20 models of (1) a Smad2 MH2 domain trimer, (2) three MAN1 UHM domains each binding to a Smad2 monomer, (3) a selected model of the complex, (4) a model of the complex in which one of the Smad2 subunits (in blue) is replaced by a Smad4 subunit (in green), (5) a zoom on the MAN1/Smad2 interface showing in sticks the side-chains of Smad2 Y366 and W386 and MAN1 Q765 and W766.
Movie presenting the crystal structure of the C-terminal region of S. cerevisiae MutLa heterodimer constituted of Mlh1 (light and dark green) and Pms1 (yellow and orange)
The movie shows successively (1) the overall structure in surface mode, (2) the C-terminus of Pms1 (yellow ribbon and stick) interacting with Mlh1 (3) the C-terminus of Mlh1 (green ribbon and stick) participating in Pms1 endonuclease site and finaly (4) interaction of a peptide (yellow) with Mlh1. The peptide contains the Mlh1 Interacting Protein motif RSK(Y/F)F, a motif used by Mlh1 to recruit Exo1, Ntg2 and Sgs1 proteins
Movie presenting a model of the secondary interaction surface of the telomeric TRF2/RAP1 assembly
The crystal structure of the dimerization domain of TRF2 is represented in yellow (1), and two model of RAP1 including residues 1 to 211 in cyan (2). The lysines involved in the interaction are coloured in blue and green, the lysines not involved in the interaction are coloured in dark grey. RAP1 specific interaction motif YxLxP is coloured in purple.